The Data Science Lab
Support Vector Machines Using Accord.NET
A support vector machine (SVM) is a software system that can perform binary classification. For example, you can use an SVM to create a model that predicts the sex of a person (male, female) based on their age, annual income, height and weight.
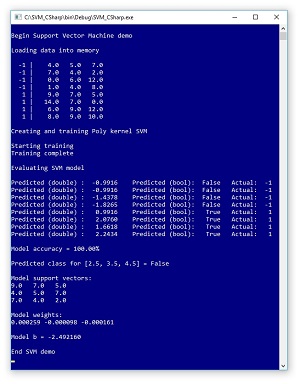
The best way to get an idea of what SVMs do is to take a look at the image of a demo program in Figure 1. The demo creates a dummy binary classification problem with just eight training items. Each item has three predictor values, and the class to predict is encoded as -1 or +1. The data is artificial but you can imagine it represents a problem where the goal is to predict if a patient will die (-1) or survive (+1) based on numeric scores of a blood test, a heart test and a lung test.
The demo program creates an SVM using a polynomial kernel. I'll explain what that means shortly. After the SVM model was trained it was used to compute predicted outputs for the training data. A computed output value of False corresponds to -1 and a computed output value of True corresponds to +1 so you can see the trained model predicted all eight training items correctly.
 [Click on image for larger view.]
Figure 1. Support Vector Machine Classification in Action
[Click on image for larger view.]
Figure 1. Support Vector Machine Classification in Action
Next, the demo computed an output value for a previously unseen input of (2.5, 3.5, 4.5) and because the computed output is False, the predicted class is whatever label is associated with -1. The demo concludes by displaying some of the trained model's information. Three of the eight input vectors are identified as special support vectors: (9, 7, 5), (4, 5, 7) and (7, 4, 2). Each support vector has an associated weight: (0.000259, -0000098, -0.000161). The SVM model has a single bias value, which is -2.49 for the demo.
This article assumes you have intermediate or better programming skill with a C-family language but doesn't assume you know anything about SVMs. The demo program is coded using C# and because it relies on the Accord.NET machine learning code library, it's not feasible to refactor to another language. However, once you understand how SVMs work, you'll find it relatively easy to use an SVM library coded in a different language such as Java or Python.
The complete source code for the demo program is presented in this article. The source code is also available in the accompanying file download. All normal error checking has been removed to keep the main ideas as clear as possible.
Understanding SVM Mechanics
SVMs are complicated. In my opinion, they're best understood by working through a concrete example. The first piece of background knowledge required is understanding what a kernel function is. Suppose you have two vectors, x1 = (6, 3, 1) and x2 = (4, 3, 5). The simplest possible kernel function is called the linear kernel and it's calculated as K(x1, x2) = (6 * 4) + (3 * 3) + (1 * 5) = 38. The pair of vectors produces a single scalar value.
The demo program uses a polynomial kernel, which is an extension of the linear kernel. If you set gamma = 1.5, degree = 2, and r = 0, then K(x1, x2) = ( 1.5 * [(6 * 4) + (3 * 3) + (1 * 5)] + 0 )^2 = 57.0^2 = 3249.
When working with SVMs, you must pick a kernel function, and supply any parameters specific to the function, such as gamma, degree, and r for the polynomial kernel. The most common kernel function used with SVMs is one called the radial basis function (RBF) kernel. The choice of kernel function and its parameters is a matter of trial and error.
The next prerequisite for understanding SVMs is the input-output mechanism. Suppose you somehow know the values of the support vectors, the weights, and the bias, as shown in Figure 1. The prediction value for the last input vector x' = (8, 9, 10) is 2.2434 and is computed as follows. You first compute the kernel function on the input vector x' with each of the three support vectors. For example, if gamma = 1, degree = 2, and r = 0 then the polynomial kernel function on x' = (8, 9, 10) and the first support vector x1 = (9, 7, 5) is K(x1, x') = 34225.0. Similarly, K(x2, x')= 21609.0 and K(x3, x') = 12544.0.
To compute the SVM output, sometimes called the decision value, you multiply each kernel result by its associated weight, sum, then add the bias value:
= (34225 * 0.000259) + (21609 * -0.000098) + (12544 * -0.000161) + (-2.492160)
= +2.2434
If the computed decision value is negative, the predicted class is -1 and if the decision value is positive, the predicted class is +1. OK, this is fine, but where do the support vectors, the weights, and the bias come from?
Support Vectors and Linear Separability
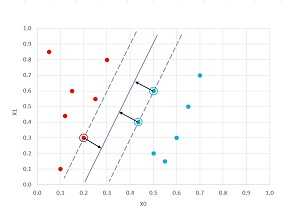
The ideas behind SVMs are shown in the graph in Figure 2. The goal of an SVM is to find the line that best separates two classes, where best separation means finding the widest gap between classes. In the graph, the one circled red dot and the two circled blue dots are the support vectors that define the best separating line.
 [Click on image for larger view.]
Figure 2. SVM Classification
[Click on image for larger view.]
Figure 2. SVM Classification
After the support vectors and best separating line have been determined, it's easy to classify a new input vector/point. There are several algorithms that can be used to determine the support vectors, weight and bias. The most common is called the sequential minimal optimization (SMO) algorithm.
The data points in Figure 2 are what's called linearly separable because there is a straight line that can separate the two classes. Unfortunately, most real-life data is not linearly separable. That's where the kernel function comes into play. Using a kernel function translates data that's not linearly separable into data which is linearly separable, and therefore can be handled by an SVM.
Program Structure
To create the demo program I launched Visual Studio and created a new console application named SVM_CSharp. I used Visual Studio 2017 with .NET Framework 4.7 but any relatively recent version of VS should work. After the template code loaded, I right-clicked on file Program.cs in the Solution Explorer window and renamed the file to SVM_Program.cs and then allowed VS to automatically rename class Program for me.
Although it's possible to write SVM code from scratch, that approach takes a lot of time and so in most situations it's better to use a code library. The demo program uses the open source Accord.NET library. In the Solution Explorer window, I right-clicked on the SVM_CSharp project name and selected the Manage NuGet Packages option. In the NuGet manager window, I selected the Browse tab and then searched for Accord.MachineLearning and when it appeared in the result list, I selected that package and then clicked the Install button.
The overall structure of the demo program, with a few minor edits to save space, is shown in Listing 1. At the top of the editor window I removed all using statements except the one referencing the top-level System namespace. Then I added two references to the Accord.NET library as shown in the code listing.
Listing 1: Overall Program Structure
using System;
using Accord.MachineLearning.VectorMachines.Learning;
using Accord.Statistics.Kernels;
namespace SVM_CSharp
{
class SVM_Program
{
static void Main(string[] args)
{
Console.WriteLine("Begin Support Vector Machine demo ");
// set up training data
// create and train SVM
// evaluate SVM model
// use model to make a new prediction
// display model information
Console.WriteLine("End SVM demo ");
Console.ReadLine();
} // Main
} // Program
} // ns
In a realistic scenario, you'd read data from a text file into memory but the eight training items in the demo are hard-coded like so:
double[][] X = {
new double[] { 4,5,7 }, new double[] { 7,4,2 },
new double[] { 0,6,12 }, new double[] { 1,4,8 },
new double[] { 9,7,5 }, new double[] { 14,7,0 },
new double[] { 6,9,12 }, new double[] { 8,9,10 } };
int[] y = { -1, -1, -1, -1, 1, 1, 1, 1 };
For simplicity and clarity, the demo does not normalize the predictor values, but in a non-demo scenario you should definitely normalize your data. The most common approach is to use min-max normalization which scales all predictor values to a [0.0, 1.0] range. After the dummy training data has been set up, it's displayed:
for (int i = 0; i < X.Length; ++i) {
Console.Write(y[i].ToString().PadLeft(4) + " | ");
for (int j = 0; j < X[i].Length; ++j) {
Console.Write(X[i][j].ToString("F1").PadLeft(6));
}
Console.WriteLine("");
}
For each line, the demo displays the class label -1/+1 value first, followed by the predictor values. One of the oldest and most widely used SVM libraries, LibSvm, requires this label-first format and so it's common to display SVM data in this way.
Creating and Training the SVM Model
The SVM model is prepared by configuring the optimization algorithm:
Console.WriteLine("Creating and training Poly kernel SVM");
var smo = new SequentialMinimalOptimization<Polynomial>();
smo.Complexity = 1.0;
smo.Kernel = new Polynomial(2, 0.0);
smo.Epsilon = 1.0e-3;
smo.Tolerance = 1.0e-2;
The Complexity property controls how complicated the decision boundary line is allowed to become. Higher values can give better accuracy at the expense of increased likelihood of model overfitting. A good value for Complexity must be determined by trial and error. The Epsilon and Tolerance properties also control the SMO algorithm. The values used, 1.0e-3 and 1.0e-2, are the default values and so I could have left out the two statements that set those properties.
The Polynomial object constructor accepts a degree argument and an r-constant argument, but does not accept a gamma argument. This effectively sets gamma to a constant value of 1.0 and is a quirk of the Accord.NET library. But the library is very well-organized and it's easy to write your own custom kernel function if you wish. The SVM model is trained like so:
Console.WriteLine("Starting training");
var svm = smo.Learn(X, y);
Console.WriteLine("Training complete");
Instead of explicitly creating an SVM object, the Learn method creates an SVM object behind the scenes, trains the SVM object, and returns the trained object.
Evaluating and Using the SVM Model
After the SVM model has been trained, it's evaluated using these statements:
Console.WriteLine("Evaluating SVM model");
bool[] preds = svm.Decide(X);
double[] score = svm.Score(X);
The Decide method returns predicted class labels as Boolean values, False for -1 and True for +1. The Score method returns predicted class labels in raw numeric form, where any negative value maps to class -1 and any positive value maps to class +1.
The demo evaluates the training data but in a non-demo scenario, you'd typically separate your original data into a training set and a test set, and evaluate the test data to get a rough approximation of how well the trained model would do on new, previously unseen data.
The demo examines each data item using these statements:
int numCorrect = 0; int numWrong = 0;
for (int i = 0; i < preds.Length; ++i) {
Console.Write("Predicted (double) : " + score[i] + " ");
Console.Write("Predicted (bool): " + preds[i] + " ");
Console.WriteLine("Actual: " + y[i]);
if (preds[i] == true && y[i] == 1) ++numCorrect;
else if (preds[i] == false && y[i] == -1) ++numCorrect;
else ++numWrong;
}
double acc = (numCorrect * 100.0) / (numCorrect + numWrong);
Console.WriteLine("Model accuracy = " + acc);
Most SVM libraries have a built-in function to compute classification accuracy. The Accord.NET library doesn't, but as you can see, the code is simple. The model is used to make a prediction like so:
bool predClass = svm.Decide(new double[] { 2.5, 3.5, 4.5 });
Console.WriteLine("Predicted class for [2.5, 3.5, 4.5] = " +
predClass);
Because the demo did not normalize the training data, you don't need to normalize when making a prediction. But in a non-demo scenario, where you normalize data, you'd need to normalize input values.
Displaying SVM Model Information
The demo displays the values of the support vectors using this code:
Console.WriteLine("Model support vectors: ");
double[][] sVectors = svm.SupportVectors;
for (int i = 0; i < sVectors.Length; ++i) {
for (int j = 0; j < sVectors[i].Length; ++j) {
Console.Write(sVectors[i][j].ToString("F1") + " ");
}
Console.WriteLine("");
}
The SupportVectors property returns an array-of-arrays style matrix. Unlike many SVM libraries, Accord.NET does not have an easy way to get the indices of the training data that correspond to the support vectors. If you have N training items, there can be as few as two support vectors or as many as N. The model weights are displayed:
Console.WriteLine("Model weights: ");
double[] wts = svm.Weights;
for (int i = 0; i < wts.Length; ++i)
Console.Write(wts[i].ToString("F6") + " ");
Console.WriteLine("");
If you read about SVMs, you have to be a bit careful because weights can be defined to either include the known y values of -1 or +1 (the usual definition) or not include the y values. The demo concludes by displaying the model bias value:
. . .
double b = svm.Threshold;
Console.WriteLine("Model b = " + b.ToString("F6"));
Console.WriteLine("End SVM demo ");
Console.ReadLine();
} // Main
} // Program
Although it's not necessary to display the weights and bias values for a trained SVM model, it can be a good way to help diagnose problems.
Wrapping Up
Although SVMs were designed to perform binary classification, many SVM libraries have extensions to perform multiclass classification and regression. In my opinion, SVMs are not well-suited for these types of problems and for them I prefer neural networks.
If you want to implement an SVM model using C# you don't have many options. Other than the Accord.NET library demonstrated in this article, the main alternative is to use a C# wrapper library around the LibSvm library. The LibSvm library is written in C++ and available as a compiled DLL so it's possible to call into the DLL using .NET Interop functionality. But, based on my experience, most of these C# wrapper libraries have significant problems, including memory leakage and null pointer issues.
The relatively new open source ML.NET library, which is backed by Microsoft, does not have SVM functionality, at least at the time I'm writing this article. But because SVMs are so important in machine learning, I expect that SVMs will be added to ML.NET relatively soon.